An international team of researchers led by the Max Planck Institute for the Science of Human History has analyzed two 3,800-year-old Y. pestis genomes that suggest a Bronze Age origin for bubonic plague.

The strain identified by the researchers was recovered from individuals in a double burial in the Samara region of Russia, who both had the same strain of the bacterium at death. The study shows that this strain is the oldest sequenced to date that contains the virulence factors considered characteristic of the bubonic plague, and is ancestral to the strains that caused the Justinian Plague, the Black Death and the 19th century plague epidemics in China.
The plague, caused by the bacterium Yersinia pestis, was the cause of some of the world’s deadliest pandemics, including the Justinian Plague, the Black Death, and the major epidemics that swept through China in the late 1800s. The disease continues to affect populations around the world today. Despite its historical and modern significance, the origin and age of the disease are not well understood. In particular, exactly when and where Y. pestis acquired the virulence profile that allows it to colonize and transmit through the flea vector has been unclear.
Recent studies of ancient Y. pestis genomes identified its earliest known variants, dating to the Late Neolithic and Early Bronze Age, but these genomes did not show the genetic signatures thought to make the plague particularly efficient – namely, adaptation to survival in fleas, which act as the main vectors that transmit the disease to mammals. This study aimed to look at more Bronze Age Y. pestis genomes, in order to investigate when and where these important adaptations occurred.
Oldest bubonic plague genome to date
In the study, the researchers analyzed nine individuals from tombs at a site in Russia. Two of the individuals were determined to have been infected with Y. pestis at the time of their deaths. The two were buried together in a single grave and were dated to approximately 3,800 years old. Analysis of the human DNA showed that the individuals were likely from the Samara-region Srubnaya-culture, which matches with the archaeological evidence. “Both individuals appear to have the same strain of Y. pestis,” comments Kirsten Bos of the Max Planck Institute for the Science of Human History. “And this strain has all the genetic components we know of that are needed for the bubonic form of the disease. So plague, with the transmission potential that we know today, has been around for much longer than we thought.”
The researchers used the data collected in combination with previously sequenced Y. pestis strains to calculate the age of their newly identified lineage at around 4,000 years. This pushes back the proposed age of the bubonic plague by 1,000 years. “Our Y. pestis isolates from around 4,000 years ago possessed all the genetic characteristics required for efficient flea transmission of plague to rodents, humans and other mammals,” states Maria Spyrou of the Max Planck Institute for the Science of Human History, first author of the study.
Early stages of the evolution of one of humankind’s most notorious pathogens
Although prior studies had identified a single lineage of Y. pestis to be present across Eurasia during the Bronze Age, the current study suggests that there were at least two plague lineages circulating contemporaneously, and that they may have encompassed different transmission and virulence characteristics. “Whether the lineages were equally prevalent in human populations, and the extent to which human activities contributed to their spread, are questions that would need further investigation,” explains senior author Johannes Krause of the Max Planck Institute for the Science of Human History. “Additional Bronze Age and Iron Age plague genomes could help pinpoint key events that contributed to the high virulence and spread of one of humankind’s most notorious pathogens,” he adds.
(Source: https://www.archaeology.wiki/blog/2018/06/11/oldest-bubonic-plague-genome-decoded/)

NovoScriptorium: Now, let’s have a brief look at recent relative research

Summary The bacteria Yersinia pestis is the etiological agent of plague and has caused human pandemics with millions of deaths in historic times. How and when it originated remains contentious. Here, we report the oldest direct evidence of Yersinia pestis identified by ancient DNA in human teeth from Asia and Europe dating from 2,800 to 5,000 years ago. By sequencing the genomes, we find that these ancient plague strains are basal to all known Yersinia pestis. We find the origins of the Yersinia pestis lineage to be at least two times older than previous estimates. We also identify a temporal sequence of genetic changes that lead to increased virulence and the emergence of the bubonic plague. Our results show that plague infection was endemic in the human populations of Eurasia at least 3,000 years before any historical recordings of pandemics.
(Source: “Early Divergent Strains of Yersinia pestis in Eurasia 5,000 Years Ago”, by Simon Rasmussen et al., 2015)

Summary The bacteria Yersinia pestis is the etiological agent of plague and has caused human pandemics with millions of deaths in historic times. How and when it originated remains contentious. Here, we report the oldest direct evidence of Yersinia pestis identified by ancient DNA in human teeth from Asia and Europe dating from 2,800 to 5,000 years ago. By sequencing the genomes, we find that these ancient plague strains are basal to all known Yersinia pestis. We find the origins of the Yersinia pestis lineage to be at least two times older than previous estimates. We also identify a temporal sequence of genetic changes that lead to increased virulence and the emergence of the bubonic plague. Our results show that plague infection was endemic in the human populations of Eurasia at least 3,000 years before any historical recordings of pandemics.
(Source: “Early Divergent Strains of Yersinia pestis in Eurasia 5,000 Years Ago”, by Simon Rasmussen et al., 2016)
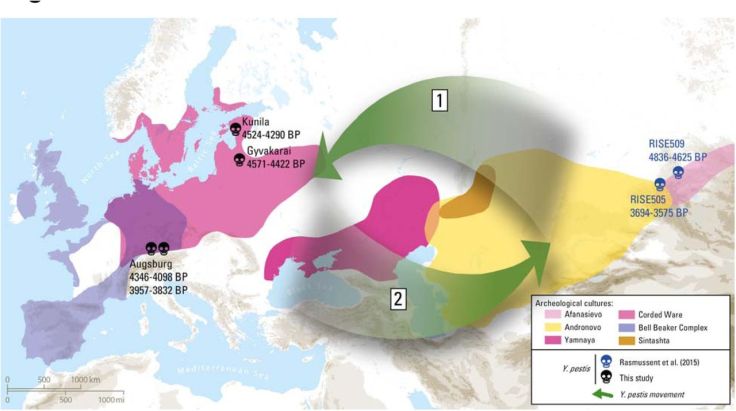
Abstract Molecular signatures of Yersinia pestis were recently identified in prehistoric Eurasian individuals, thus suggesting Y. pestis caused some form of disease in humans prior to the first historically documented pandemic. Here, we present six new Y. pestis genomes spanning from the European Late Neolithic to the Bronze Age (LNBA) dating from 4,800 to 3,700 BP. We show that all currently investigated LNBA strains form a single genetic clade in the Y. pestis phylogeny that appears to be extinct. Interpreting our data within the context of recent ancient human genomic evidence, which suggests an increase in human mobility during the LNBA, we propose a possible scenario for the spread of Y. pestis during the LNBA: Y. pestis may have entered Europe from Central Eurasia during an expansion of steppe people, persisted within Europe until the mid Bronze Age, and moved back towards Central Eurasia in parallel with subsequent human population movements.
(Source: “The Stone Age Plague: 1000 years of Persistence in Eurasia”, by Aida Andrades Valtueña et al., 2017)
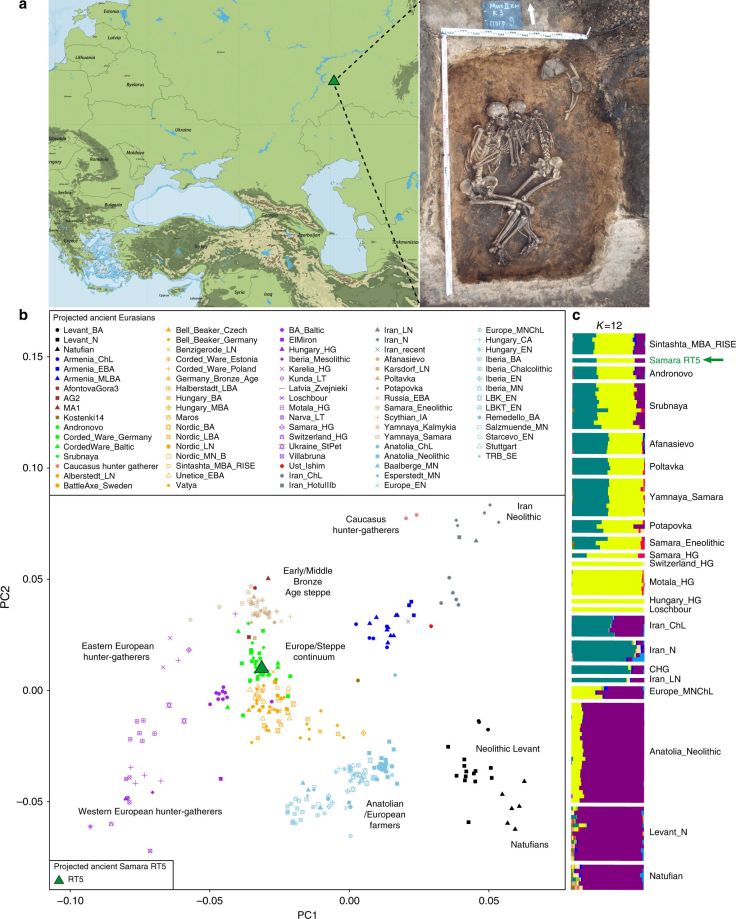

SummaryYersinia pestis, the etiologic agent of plague, is a bacterium associated with wild rodents and their fleas. Historically it was responsible for three pandemics: the Plague of Justinian in the 6th century AD, which persisted until the 8th century; the renowned Black Death of the 14th century, with recurrent outbreaks until the 18th century; and the most recent 19th century pandemic, in which Y. pestis spread worldwide and became endemic in several regions. The discovery of molecular signatures of Y. pestis in prehistoric Eurasian individuals and two genomes from Southern Siberia suggest that Y. pestis caused some form of disease in humans prior to the first historically documented pandemic. Here, we present six new European Y. pestis genomes spanning the Late Neolithic to the Bronze Age (LNBA; 4,800 to 3,700 calibrated years before present). This time period is characterized by major transformative cultural and social changes that led to cross-European networks of contact and exchange. We show that all known LNBA strains form a single putatively extinct clade in the Y. pestis phylogeny. Interpreting our data within the context of recent ancient human genomic evidence that suggests an increase in human mobility during the LNBA, we propose a possible scenario for the early spread of Y. pestis: the pathogen may have entered Europe from Central Eurasia following an expansion of people from the steppe, persisted within Europe until the mid-Bronze Age, and moved back toward Central Eurasia in parallel with human populations.
(Source: “The Stone Age Plague and Its Persistence in Eurasia”, by Aida Andrades Valtueña et al., 2017)
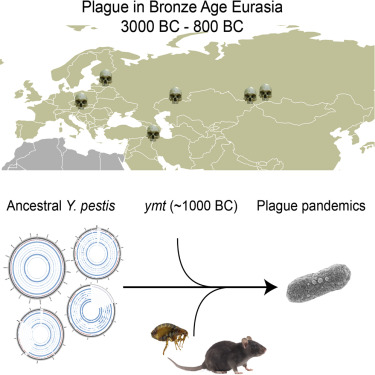
NovoScriptorium: Now, let’s take a closer look at some extracts from the published work mentioned in the sourced article

Abstract The origin of Yersinia pestis and the early stages of its evolution are fundamental subjects of investigation given its high virulence and mortality that resulted from past pandemics. Although the earliest evidence of Y. pestis infections in humans has been identified in Late Neolithic/Bronze Age Eurasia (LNBA 5000–3500y BP), these strains lack key genetic components required for flea adaptation, thus making their mode of transmission and disease presentation in humans unclear. Here, we reconstruct ancient Y. pestis genomes from individuals associated with the Late Bronze Age period (~3800 BP) in the Samara region of modern-day Russia. We show clear distinctions between our new strains and the LNBA lineage, and suggest that the full ability for flea-mediated transmission causing bubonic plague evolved more than 1000 years earlier than previously suggested. Finally, we propose that several Y. pestis lineages were established during the Bronze Age, some of which persist to the present day.

IntroductionYersinia pestis, the causative agent of bubonic, pneumonic and septicaemic plague, evolved from the closely related environmental progenitor Y. pseudotuberculosis. Although primarily a coloniser of sylvatic rodents via flea-dependent transmission, ancient DNA studies have demonstrated its status as an infectious disease in humans for the last 5000 years, and have confirmed its involvement in some of the most devastating historical pandemics. The first historically recorded plague pandemic began with the Plague of Justinian (AD 541–543), and persisted until the eighth century AD. The second pandemic occurred between the 14th and 18th centuries AD, began with the infamous Black Death of Europe in 1347 and was a precursor of modern-day plague epidemics over a wide geographic range.
After its divergence from Y. pseudotuberculosis, Y. pestis acquired its high pathogenicity and distinct niche mainly by chromosomal gene loss as well as the acquisition of two virulence-associated plasmids, pMT1 and pPCP. Throughout this process, one of the most crucial evolutionary adaptations related to its pathogenicity was its ability to colonise arthropods, a phenotypic/functional gain mediated by a combination of chromosomal and plasmid loci. These genetic changes are central to the most common “bubonic” form of the disease, where bacteria enter the body via the bite of an infected flea, travel via the lymph to the closest lymph node and replicate while evading host defences. Recent ancient genomic investigations of Y. pestis have identified its earliest known variants in Eurasia during the Late Neolithic/Bronze Age period (LNBA) that show genetic characteristics incompatible with arthropod adaptation. These strains, therefore, have been considered incapable of an efficient flea-based transmission; however, the alternative early-phase transmission could have provided an independent means of arthropod dissemination. To date, the earliest evidence of a Y. pestis strain with signatures associated with flea adaptation has been reported during the Iron Age through shotgun sequencing of an ~2900-year-old genome from Armenia (strain RISE397), though at a coverage too low (0.25-fold) to permit confident phylogenetic positioning. Although the mechanism by which the LNBA lineage caused human disease is unclear, its frequency in Eurasia during the Bronze Age and its phylogeographic pattern that mimics contemporaneous human migrations are noteworthy.
The central steppe region seems to have played a significant role as a migration corridor during the entire Bronze Age, and as such, it likely facilitated the spread of human-associated pathogens, such as Y. pestis, across Eurasia. Here, we explore additional Y. pestis diversity in that region by isolating strains from LBA Samara, in Russia. We identify a Y. pestis lineage contemporaneous to the LNBA strains with genomic variants consistent with flea adaptation. This reveals the co-circulation of two Y. pestis lineages during the Bronze Age with different properties in terms of their transmission and disease potentials.
Results
Y. pestis and human endogenous DNA screening
We screened a total of 64 million shotgun next-generation sequencing (NGS) reads from nine teeth of nine individuals recovered from kurgan burials in the Samara region (see Supplementary Methods) to assess the endogenous human DNA content and the possible presence of Y. pestis. Our screening revealed four potentially positive individuals, one of which, individual RT5, exhibited the highest amounts of endogenous Y. pestis DNA.
Y. pestis virulence factor analysis
The presence of several Y. pestis virulence-associated genes was evaluated in RT5. While the LNBA, 0.PE2 and 0.PE4 strains seem to lack certain virulence determinants, RT5 harbours all known virulence factors with the exception of the filamentous prophage (YpfΦ), which is, however, most consistently identified among 1.ORI strains.
Another important gene for Y. pestis virulence is pla located on the species-specific pPCP1 plasmid. Although the gene is largely conserved among Y. pestis strains, an isoleucine (ancestral) to threonine (derived) alteration at amino acid position 259 has been used to differentiate the most basal isolates (LNBA, 0.PE7, 0.PE2 and 0.PE4) from the rest of Y. pestis. In RT5, the pla genotype was manually explored, and was found to exhibit the ancestral allele. Although the ancestral pla allele has been associated with a less-efficient bacterial dissemination in mammals, modern strains from lineages 0.PE4 and 0.PE7 have proven to be potent inducers of bubonic plague in humans.

Discussion We identify two ~3800-year-old individuals out of nine analysed that were infected with Y. pestis at the time of their deaths. The Samara Y. pestis genomes presented here reveal greater lineage diversity during the Bronze Age than was previously described.
Our dating analyses consistently suggest the presence of this putative ancestor at ~4000y BP followed by a population expansion shortly after that time. Though its place of origin is not yet empirically identified, given the close genetic and temporal affinity to RT5, a steppe source is plausible. Given that previous research has proposed a relationship between rapid Y. pestis expansions and historical plague epidemics in humans, future investigations of lineage diversity from modern and ancient sources may reveal additional details on this ancient radiation event.
A recent study has suggested that flea-adapted Y. pestis, along with its potential to cause bubonic plague in humans, likely originated around 3000y BP. Contrary to such conclusions, the lineage giving rise to our Y. pestis isolates (RT5 and RT6) likely arose ~4000 years ago, and possessed all vital genetic characteristics required for flea-borne transmission of plague in rodents, humans and other mammals.
Overall, the detection of Y. pestis in Bronze Age human remains from Eurasia has suggested the presence of the pathogen in this vast geographic area along with its ability to cause bubonic plague millennia before the first historically documented plague pandemic. It seems possible that already in the Bronze Age, with the establishment of transport and trade networks, the interconnectivity between Europe and Asia that is also reflected in the ancient human genomes, likely contributed to the spread of infectious disease. Similarly, the abundant trade routes of the medieval period are considered the main conduit for plague’s movement between Asia and Europe. Our current data suggest a more complex model, where at least two human-associated lineages (LNBA and RT5) with different transmission potentials were established in Eurasia during the Bronze Age.
Additional Bronze Age/Iron Age genomes could provide further insights into the early stages of Y. pestis evolution, and help pinpoint key events that contributed to the high virulence and spread of one of humankind’s most notorious pathogens.
(Source: “Analysis of 3800-year-old Yersinia pestis genomes suggests Bronze Age origin for bubonic plague”, by Maria A. Spyrou et al., 2018)

Research-Selection for NovoScriptorium: Maximus E. Niles

Leave a comment